Circulating recombinant forms of HBV identified with jpHMM
For each circulating recombinant form identified with jpHMM (represented by at least two unrelated sequences) the predicted recombination of one reference strain is given.
Please note that only recombinant forms that do not contain any uncertainty regions are given.
In addition to the recombinant forms presented here unique HBV recombinant forms that are represented by only one single sequence have been identified with jpHMM. They can be found here: unique HBV recombinant forms
Please click on an image to zoom into it.
A link to the list of the Genbank accession numbers of all sequences
clustered with the given representative recombinant strain can be opened by a click on the given Genbank accession number.
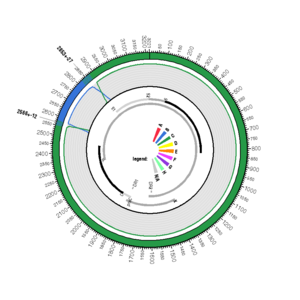 EU796069
EU796069
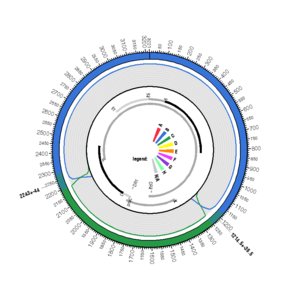 EU939627
EU939627
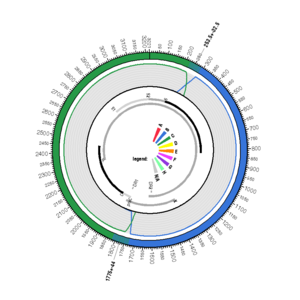 EU939628
EU939628
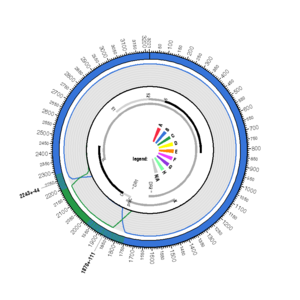 D00330
D00330
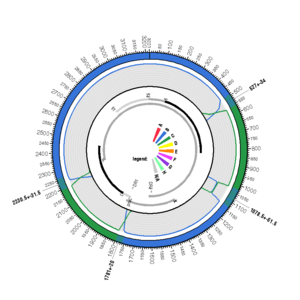 FJ386674
FJ386674
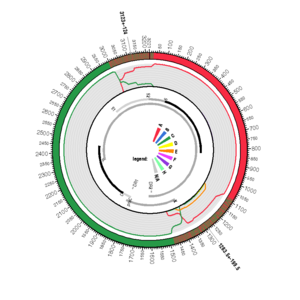 AB231908
AB231908
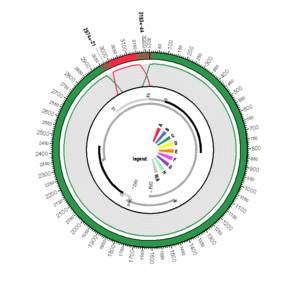 AY057947
AY057947
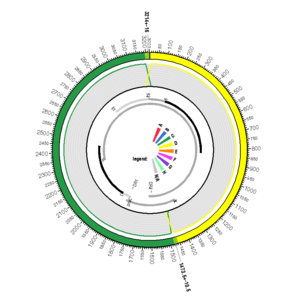 AY057948
AY057948
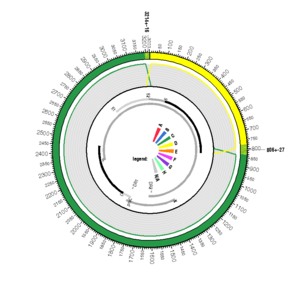 AF461043
AF461043
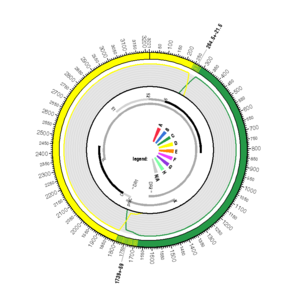 FJ562223
FJ562223
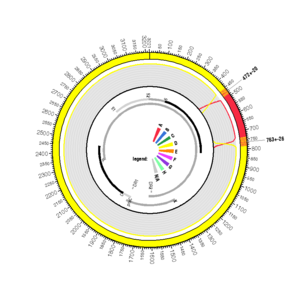 AF418681
AF418681
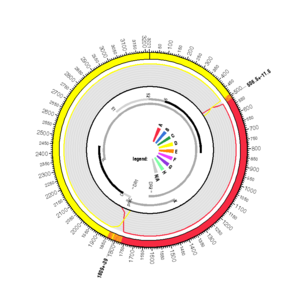 AY236161
AY236161
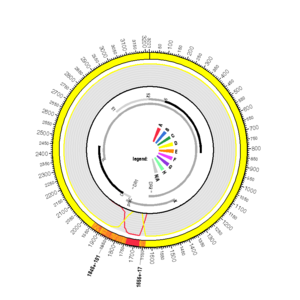 X80925
X80925
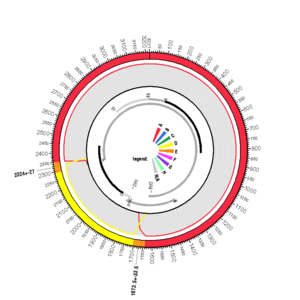 AF418674
AF418674
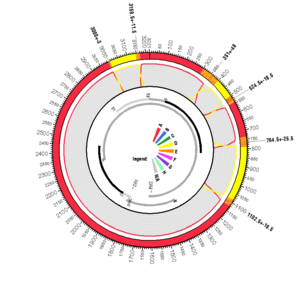 AF418685
AF418685
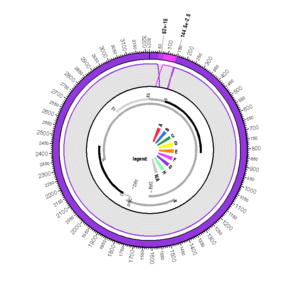 EF464097
EF464097
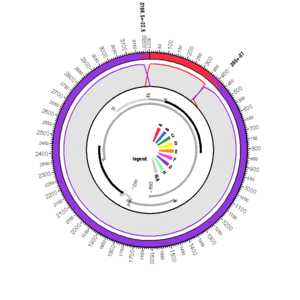 EF464099
EF464099
All images were created with the software package Circos (Krzywinski, M. et al., Genome Res. (2009)).
References
Please cite one of the following papers if you use this tool in your publication (a list of all references can be found here):
- A.-K. Schultz, M. Zhang, I. Bulla, T. Leitner, B. Korber, B. Morgenstern, M. Stanke. jpHMM: Improving the reliability of recombination prediction in HIV-1. Nucleic Acids Research, 37:W647-51. 2009
- M. Zhang, A.-K. Schultz, C. Calef, C. Kuiken, T. Leitner, B. Korber, B. Morgenstern, M. Stanke. jpHMM at GOBICS: a web server to detect genomic recombinations in HIV-1. Nucleic Acids Research, 34:W463-5. 2006.
- A.-K. Schultz, M. Zhang, T. Leitner, C. Kuiken, B. Korber, B. Morgenstern, M. Stanke. A Jumping Profile Hidden Markov Model and Applications to Recombination Sites in HIV and HCV Genomes. BMC Bioinformatics 7:265. 2006.
Questions or comments? Email contact
Copyright � 2005-2006 Dept. of Bioinformatics (IMG)